Ten Jupyter/IPython essentials
In this section, we will cover ten essential features of Jupyter and IPython that make them so useful for interactive computing.
Using IPython as an extended shell
Note
Unfortunately, this subsection will not work well on Windows. The goal here is to demonstrate accessing the operating system's shell from IPython. We could say that, by design, the Windows shell is much more limited than those provided by Linux and OS X. Windows favors user interactions from the graphical interface, whereas Linux and OS X inherit Unix's flexible command-line capabilities. If you want to share and distribute your notebooks, you shouldn't rely on the techniques exposed in this subsection. Rather, you should use the Python equivalents, which are more verbose but also more powerful. Using the shell from IPython is only useful during interactive sessions of users already familiar with the Unix shell.
Open a terminal and type the following commands to go to the minibook's chapter1 directory and launch the Notebook server:
$ cd ~/minibook/chapter1/ $ jupyter notebook
In the Notebook dashboard, open the 15-ten.ipynb notebook. You can also create a new notebook if you prefer not to use the book's code.
Let's illustrate how to use IPython as an extended shell. We will download an example dataset, navigate through the filesystem, and open text files, all from the Notebook. The dataset contains social network data of hundreds of volunteer Facebook users. This BSD-licensed dataset is provided freely by Stanford's SNAP project (http://snap.stanford.edu/data/).
IPython provides several magic commands that let you interact with your filesystem. These commands are prefixed with a %. For example here is how to display the current working directory:
In [1]: %pwd Out[1]: '/home/cyrille/minibook/chapter1'
Note
Like most other magic commands, this magic command works on all operating systems, including Windows. IPython implements several cross-platform Python equivalents of common Unix commands like pwd. For other commands not implemented by IPython, we need to call shell commands directly with the ! prefix (as shown in the following examples). This doesn't work well on Windows since many of these commands are Unix-specific. In brief, %-prefixed commands should work on all operating systems while !-prefixed commands will generally only work on Linux and OS X, not Windows.
Let's download the dataset from the book's data repository (https://github.com/ipython-books/minibook-2nd-data). IPython doesn't yet provide a magic command for downloading data, but we can use another IPython trick: we can run any system or terminal command from IPython by prefixing it with an exclamation mark (!). For example, here is how to use the wget download utility only available on Unix systems:
In [2]: !wget https://raw.githubusercontent.com/ipython-books/minibook-2nd-data/master/facebook.zip
Note
If wget is not installed, you can install it with your OS package manager. For example, on Ubuntu: sudo apt-get install wget; on OS X: brew install wget. On OS X, brew is available at http://brew.sh/. On Windows, you should download the file manually from the data repository, as explained later.
This wget command downloads a file from a URL and saves it to a file in the local filesystem. Let's display the list of files in the current directory using the %ls magic command (available on all systems, even on Windows, since it is a magic command provided by IPython), as follows:
In [3]: %ls Out[3]: facebook.zip [...]
We see a new facebook.zip file.
Note
If you are on Windows, or if downloading the file from IPython didn't work, you can always download this file manually via your web browser at the following URL: https://github.com/ipython-books/minibook-2nd-data/. Then save the Facebook dataset in the current directory (the one containing this notebook, which should be ~/minibook/chapter1/).
The next step is to unzip this file in the current directory. The first way of doing it is to use your operating system, generally with a right-click on the icon. On Linux and OS X, we can also use the unzip command-line tool (you may need to install it first, for example with a command like sudo apt-get install unzip on Ubuntu). Finally, it is also possible to do it in pure Python with the zipfile module (see https://docs.python.org/3.4/library/zipfile.html).
Here, we'll call the unzip tool, which will only work on Linux and OS X, not Windows:
In [4]: !unzip facebook.zip
Once the archive has been extracted, a new subdirectory named facebook appears, as shown here:
In [5]: %ls Out[5]: facebook facebook.zip [...]
Let's enter into this subdirectory with the %cd magic command (all operating systems), as follows:
In [6]: %cd facebook Out[6]: /home/cyrille/minibook/chapter1/facebook
IPython provides a %bookmark magic to create an alias to the current directory. Let's type the following:
In [7]: %bookmark fbdata
Now, in any future session, we'll be able to just type %cd fbdata to enter into this directory. Type %bookmark? to see all options. This magic command is helpful when dealing with many directories.
Let's display the contents of the directory:
In [8]: %ls Out[8]: 0.circles 1684.circles 3437.circles 3980.circles 686.circles 0.edges 1684.edges 3437.edges 3980.edges 686.edges 107.circles 1912.circles 348.circles 414.circles 698.circles 107.edges 1912.edges 348.edges 414.edges 698.edges
Here, every number identifies a Facebook user (called the ego user). The .edges file contains its social graph. In this graph, nodes represent other Facebook users, and edges represent friendship links between them. The .circles file contains lists of friends.
Let's retrieve the list of .edges files with the following command (which won't work on Windows):
In [9]: files = !ls -1 -S | grep .edges
The Unix command ls -1 -S lists all files in the current directory, sorted by decreasing size. The pipe | grep edges filters only those files that contain .edges. Then, this list is assigned to a new Python variable named files, as follows:
In [10]: files Out[10]: ['1912.edges', '107.edges', '1684.edges', '3437.edges', '348.edges', '0.edges', '414.edges', '686.edges', '698.edges', '3980.edges']
On Windows, you can use the following Python code to obtain the same list (if you're not on Windows, you can skip this code listing):
In [11]: import os from operator import itemgetter # Get the name and file size of all .edges files. files = [(file, os.stat(file).st_size) for file in os.listdir('.') if file.endswith('.edges')] # Sort the list with the second item (file size), # in decreasing order. files = sorted(files, key=itemgetter(1), reverse=True) # Only keep the first item (file name), in the same order. files = [file for (file, size) in files]
Let's display the first few lines of the first file in the list (Unix-specific command):
In [12]: !head -n5 {files[0]} Out[12]: 2290 2363 2346 2025 2140 2428 2201 2506 2425 2557
The curly braces {} let us insert a Python variable within a system command (here, the head Unix command which displays the first lines of a text file).
In an .edges file, every line contains the two nodes forming every edge. The .circles file contains lists of friends. Every line contains a space-separated list of the users forming every circle.
Tip
Alias commands
If you use a complex command regularly, you can create an alias with the %alias magic command. Type %alias? for more information. See also the related %store magic command.
Learning magic commands
Besides the filesystem commands we have seen in the previous section, IPython provides many other magic commands. You can display the list of all magic commands with the %lsmagic magic command, as follows:
In [13]: %lsmagic Out[13]: Available line magics: %alias %alias_magic %autocall %automagic %autosave %bookmark %cat %cd %clear %colors %config %connect_info %cp %debug %dhist %dirs %doctest_mode %ed %edit %env %gui %hist %history %install_default_config %install_ext %install_profiles %killbgscripts %ldir %less %lf %lk %ll %load %load_ext %loadpy %logoff %logon %logstart %logstate %logstop %ls %lsmagic %lx %macro %magic %man %matplotlib %mkdir %more %mv %notebook %page %pastebin %pdb %pdef %pdoc %pfile %pinfo %pinfo2 %popd %pprint %precision %profile %prun %psearch %psource %pushd %pwd %pycat %pylab %qtconsole %quickref %recall %rehashx %reload_ext %rep %rerun %reset %reset_selective %rm %rmdir %run %save %sc %set_env %store %sx %system %tb %time %timeit %unalias %unload_ext %who %who_ls %whos %xdel %xmode Available cell magics: %%! %%HTML %%SVG %%bash %%capture %%debug %%file %%html %%javascript %%latex %%perl %%prun %%pypy %%python %%python2 %%python3 %%ruby %%script %%sh %%svg %%sx %%system %%time %%timeit %%writefile Automagic is ON, % prefix IS NOT needed for line magics.
To obtain information about a magic command, append a question mark (?) after the command, as shown in the following example:
In [14]: %history?
The %history magic command lets you display and manipulate your command history in IPython. For example, the following command shows your last five commands:
In [15]: %history -l 5 Out[15]: files = !ls -1 -S | grep .edges files !head -n5 {files[0]} %lsmagic %history?
Let's also mention the %dhist magic command that shows you a history of all visited directories.
Another useful magic command is %paste, which lets you copy-paste Python code from anywhere into the IPython console (it is not available in the Notebook, where you can copy-paste as usual).
In IPython, the underscore (_) character always contains the last output. This is useful if you ran some command and forgot to assign the output to a variable.
In [16]: # how many minutes in a day? 24 * 60 Out[16]: 1440 In [17]: # and in a year? _ * 365 Out[17]: 525600
We will now see several cell magics, which are magic commands that apply to a whole code cell rather than just a line of code. They are prefixed by two percent signs (%%).
The %%capture cell magic lets you capture the standard output and error output of some code into a Python variable. Here is an example (the outputs are captured in the output Python variable):
In [18]: %%capture output %ls In [19]: output.stdout Out[19]: 0.circles 1684.circles 3437.circles 3980.circles 686.circles 0.edges 1684.edges 3437.edges 3980.edges 686.edges 107.circles 1912.circles 348.circles 414.circles 698.circles 107.edges 1912.edges 348.edges 414.edges 698.edges
The %%bash cell magic is an extension of the ! shell prefix. It lets you run multiline bash code in the Notebook, as shown here:
In [20]: %%bash cd .. touch _HEY ls rm _HEY cd facebook Out[20]: _HEY facebook facebook.zip [...]
More generally, the %%script cell magic lets you execute code with any program installed on your system. For example, assuming Haskell is installed (see https://www.haskell.org/downloads), you can easily execute Haskell code from the Notebook, as follows:
In [21]: %%script ghci putStrLn "Hello world!" Out[21]: GHCi, version 7.6.3: http://www.haskell.org/ghc/ :? for help Loading package ghc-prim ... linking ... done. Loading package integer-gmp ... linking ... done. Loading package base ... linking ... done. Prelude> Hello world! Prelude> Leaving GHCi.
The ghci executable runs in a separate process, and the contents of the cell are passed to the executable's input. You can also put a full path after %%script, for example, on Linux: %%script /usr/bin/ghci.
Tip
IHaskell kernel
This way of calling external scripts is only useful for quick interactive experiments. If you want to run Haskell notebooks, you can use the IHaskell notebook for Jupyter, available at https://github.com/gibiansky/IHaskell.
Finally, the %%writefile cell magic lets you write some text in a new file, as shown here:
In [22]: %%writefile myfile.txt Hello world! Out[22]: Writing myfile.txt In [23]: !more myfile.txt Out[23]: Hello world!
Now, let's delete the file, as follows:
In [24]: !rm myfile.txt
Note
On Windows, you need to type !del myfile.txt instead.
There are many other magic commands available. We will see several of them later in this book. Also, in Chapter 6, Customizing IPython, we will see how to create new magic commands. This is much easier than it sounds!
Refer to the following page for up-to-date documentation about all magic commands: http://www.ipython.org/ipython-doc/dev/interactive/magics.html.
Mastering tab completion
Tab completion is an incredibly useful feature in Jupyter and IPython. When you start to write something and press the Tab key on your keyboard, IPython can guess what you're trying to do, and propose a list of options that match what you have typed so far. This works for Python functions, variables, magic commands, files, and more.
Let's first make sure we are in the facebook directory (using the directory alias created previously):
In [25]: %cd fbdata %ls Out[25]: (bookmark:fbdata) -> /home/cyrille/minibook/chapter1/facebook /home/cyrille/minibook/chapter1/facebook 0.circles 1684.circles 3437.circles 3980.circles 686.circles 0.edges 1684.edges 3437.edges 3980.edges 686.edges 107.circles 1912.circles 348.circles 414.circles 698.circles 107.edges 1912.edges 348.edges 414.edges 698.edges
Now, start typing a command and press Tab before finishing it (here, press the Tab key on your keyboard right after typing e), as follows:
!head -n5 107.e<TAB>
IPython automatically completes the command and adds the four remaining characters (dges). IPython recognized the beginning of a file name and completed the command. If there are several completion possibilities, IPython doesn't complete anything, but instead shows a list of all options. You can then choose the appropriate solution by pressing the Up or Down keys on the keyboard, and pressing Tab again. The following screenshot shows an example:

Tab completion in the Notebook
Tab completion is extremely useful when you're getting acquainted with a new Python package. For example, to quickly see all functions provided by the NetworkX package, you can type import networkx; networkx.<TAB>.
Tip
Customizing tab completion
If you're writing a Python library, you probably want to write tab-completion-aware code. Your users who work with IPython will thank you! In most cases, you have nothing to do, and tab completion will just work. In the rare cases where you use advanced dynamic techniques in a class, you can customize tab completion by implementing a __dir__(self) method that returns all attributes available in the current class instance. See this reference for more details: https://docs.python.org/3.4/library/functions.html#dir.
Writing interactive documents in the Notebook with Markdown
You can write code and text in the Notebook. Every cell is either a Markdown cell or a code cell. The Markdown cell lets you write text. Markdown is a text formatting syntax that supports headers, bold, italics, hypertext links, images, and code. In the Notebook, you can also write mathematical equations in a Markdown cell using LaTeX, a markup language widely used for equations. Finally, you can also write some HTML in a Markdown cell, and it will be interpreted correctly.
Here is an example of a paragraph in Markdown:
### New paragraph This is *rich* **text** with [links](http://ipython.org), equations: $$\hat{f}(\xi) = \int_{-\infty}^{+\infty} f(x)\, \mathrm{e}^{-i \xi x} dx$$ code with syntax highlighting: ```python print("Hello world!") ``` and images: 
If you write this in a Markdown cell, and "play" the cell (for example, by pressing Ctrl + Enter), you will see the rendered text. The following screenshot shows the two modes of the cell:

A Markdown cell in the Notebook
By using both Markdown cells and code cells in a notebook, you can write an interactive document about any technical topic. Hence, the Notebook is not only an interface to code, it is also a platform to write documents or even books. In fact, this very book is entirely written in the Notebook!
Here are a few references about Markdown and LaTeX:
- Markdown on Wikipedia at http://en.wikipedia.org/wiki/Markdown
- The original specification, at http://daringfireball.net/projects/markdown/
- A Markdown tutorial by GitHub, at https://help.github.com/articles/markdown-basics/
- CommonMark, a standardized version of Markdown, at http://commonmark.org/
- LaTeX on Wikipedia at http://en.wikipedia.org/wiki/LaTeX
Creating interactive widgets in the Notebook
You can add interactive graphical elements called widgets in a notebook. Examples of rich graphical widgets include buttons, sliders, dropdown menus, interactive plots, as well as videos, audio files, and complete Graphical User Interfaces (GUIs). Widget support in Jupyter is still relatively experimental at this point, but we will use them at several occasions in this book. This section shows a few basic examples.
First, let's add a YouTube video in a notebook, as follows:
In [26]: from IPython.display import YouTubeVideo YouTubeVideo('j9YpkSX7NNM')
Following is a screenshot of a YouTube video in a notebook:

Youtube in the Notebook
The YoutubeVideo constructor accepts a YouTube identifier as input.
Next, let's show how to create a graphical control to manipulate the inputs to a Python function:
In [27]: from ipywidgets import interact # IPython.html.widgets before # IPython 4.0 @interact(x=(0, 10)) def square(x): print("The square of %d is %d." % (x, x**2)) Out[27]: 'The square of 7 is 49.'
Here is a screenshot:
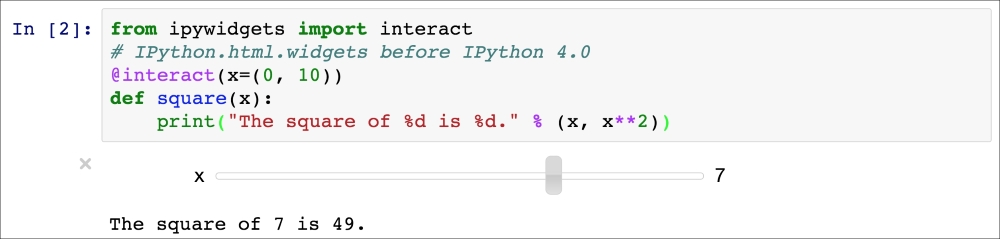
Interactive widget in the Notebook
The square(x) function just prints a sentence like The square of 7 is 49. By adding the @interact decorator above the function's definition, we tell IPython to create a widget to control the function's input x. The argument x=(0, 10) is a convention to indicate that we want a slider to control an integer between 0 and 10.
This method supports other common controls like checkboxes, dropdown menus, radio buttons, push buttons, and others.
Finally, entirely customizable widgets can be created, but this requires some knowledge of web technologies such as HTML, CSS, and JavaScript. The IPython Cookbook (http://ipython-books.github.io/cookbook/) contains many examples. You can also refer to the following links for more information:
- IPython widgets tutorial at https://github.com/ipython/ipywidgets/blob/master/examples/Index.ipynb
- Introducing the interactive features of the IPython Notebook, at https://github.com/rossant/euroscipy2014
- A piano in the Notebook, at http://nbviewer.ipython.org/github/ipython-books/cookbook-code/blob/master/notebooks/chapter03_notebook/05_basic_widgets.ipynb
Note
Most of these references describe APIs that were introduced in IPython 3.0, but are still experimental at this point. They may not work with future versions of Jupyter and IPython.
Running Python scripts from IPython
Notebooks are mainly designed for interactive exploration, not for reusability. It is currently difficult to reuse parts of a notebook in another script or notebook. Many users just copy-paste their code, which goes against the Don't Repeat Yourself (DRY) principle.
A common practice is to put frequently used code into a Python script, for example myscript.py. Such a script can be called from the system terminal like this: python myscript.py. Python will execute the script and quit at the end. If you use the -i option, Python will start the interactive prompt when the script ends.
IPython also supports this technique; just replace python by ipython. For example: ipython -i script.py to run script.py interactively with IPython.
You can also run a script from within IPython by using the %run magic command. The script runs in an empty namespace, meaning that any variable defined in the interactive namespace is not available within the executed script. However, at the end of the execution, the control returns to IPython, and the variables defined in the script are imported into the interactive namespace. This lets you inspect the intermediate variables used in the script. If you use the -i option, the script will run in the interactive namespace. Any variable defined in the interactive session will be available in the script.
Let's also mention the similar %load magic command.
Note
A namespace is a dictionary mapping variable names to Python objects. The global namespace contains global variables, whereas the local namespace of a function contains the local variables defined in the function. In IPython, the interactive namespace contains all objects defined and imported within the current interactive session. The %who, %whos, and %who_ls magic commands give you some information about the interactive variables.
For example, let's write a script egos.py that lists all ego identifiers in the Facebook data folder. Since each filename is of the form <egoid>.<extension>, we list all files, remove the extensions, and take the sorted list of all unique identifiers. We can create this file from the Notebook, using the %%writefile cell magic as follows:
In [28]: %cd fbdata %cd .. Out[28]: (bookmark:fbdata) -> /home/cyrille/minibook/chapter1/facebook /home/cyrille/minibook/chapter1/facebook In [29]: %%writefile egos.py import sys import os # We retrieve the folder as the first positional argument # to the command-line call if len(sys.argv) > 1: folder = sys.argv[1] # We list all files in the specified folder files = os.listdir(folder) # ids contains the list of idenfitiers identifiers = [int(file.split('.')[0]) for file in files] # Finally, we remove duplicates with set(), and sort the list # with sorted(). ids = sorted(set(identifiers)) Out[29]: Overwriting egos.py
This script accepts an argument folder as an input. It is retrieved from the Python script via the sys.argv list, which contains the list of arguments passed to the script via the command-line interface.
Let's execute this script in IPython using the %run magic command, as follows:
In [30]: %run egos.py facebook
Note
If you get an error when running this script, make sure that the facebook directory only contains <number>.xxx files (like 0.circles or 1684.edges).
In [31]: ids Out[31]: [0, 107, 348, 414, 686, 698, 1684, 1912, 3437, 3980]
The ids variable created in the script is now available in the interactive namespace.
Let's see what happens if we do not specify the folder name to the script, as follows:
In [32]: folder = 'facebook' In [33]: %run egos.py
We get an error: NameError: name 'folder' is not defined. This is because the variable folder is defined in the interactive namespace, but is not available within the script by default. We can change this behavior with the -i option, as follows:
In [34]: %run -i egos.py In [35]: ids Out[35]: [0, 107, 348, 414, 686, 698, 1684, 1912, 3437, 3980]
This time, the script correctly used the folder variable.
Introspecting Python objects
IPython can display detailed information about any Python object.
First, type ? after a variable name to get some information about it. For example, let's inspect NetworkX's Graph class, as follows:
In [36]: import networkx In [37]: networkx.Graph?
This shows the docstring and other information in the Notebook pager, as shown in the following screenshot:

Typing ?? instead of ? shows even more information, including the whole source code of the Python object when it is available.
There are also several magic commands for inspecting Python objects:
%pdef: Displays a function definition%pdoc: Displays the docstring of a Python object%psource: Displays the source code of an object (function, class, or method)%pfile: Displays the source code of the Python script where an object is defined
Debugging Python code
IPython makes it convenient to debug a script or an entire application. It provides interactive access to an enhanced version of the Python debugger.
First, when you encounter an exception, you can immediately use the %debug magic command to launch the IPython debugger at the exact point where the exception was raised.
If you activate the %pdb magic command, the debugger will automatically start at the very next exception. You can also start IPython with ipython --pdb.
Finally, you can run a whole script under the control of the debugger with the %run -d command. This command executes the specified script with a break point at the first line so that you can precisely control the execution flow of the script. You can also specify explicitly where to put the first breakpoint; type %run -d -b29 script.py to pause the program execution on line 29 of script.py. In all cases, you first need to type c to start the script execution.
When the debugger starts, you enter into a special prompt, as indicated by ipdb>. The program execution is then paused at a given point in the code. You can type w to display the line and stack location where the debugger has paused. At this point, you have access to all local variables and you can precisely control how you want to resume the execution. Within the debugger, several commands are available to navigate into the traceback; they are as follows:
u/dfor going up/down into the call stacksto step into the next statementnto continue execution until the next line in the current functionrto continue execution until the current function returnscto continue execution until the next breakpoint or exception
Other useful commands include:
pto evaluate and print any expressionato obtain the arguments of the current functions- The
!prefix to execute any Python command within the debugger
The entire list of commands can be found in the documentation of the pdb module in Python at https://docs.python.org/3.4/library/pdb.html.
Let's also mention the IPython.embed() function that you can call anywhere in a Python script. This stops the script execution and starts IPython for debugging purposes. Leaving the embedded IPython terminal resumes the normal execution of the script.
Benchmarking Python code
The %timeit magic function lets us estimate the execution time of any Python statement. Under the hood, it uses Python's native timeit module.
In the following example, we first load an ego graph from our Facebook dataset using the NetworkX package. Then we evaluate how much time it takes to tell whether the graph is connected or not:
Let's go to the data directory, as follows:
In [38]: %cd fbdata Out[38]: (bookmark:fbdata) -> /home/cyrille/minibook/chapter1/facebook /home/cyrille/minibook/chapter1/facebook
We load NetworkX, as follows:
In [39]: import networkx
We can load a graph using the read_edgelist() function, as follows:
In [40]: graph = networkx.read_edgelist('107.edges')
How big is our graph?
In [41]: len(graph.nodes()), len(graph.edges()) Out[41]: (1034, 26749)
Now let's find out whether the graph is connected or not:
In [42]: networkx.is_connected(graph) Out[42]: True
How long did this call take?
In [43]: %timeit networkx.is_connected(graph) Out[43]: 100 loops, best of 3: 5.92 ms per loop
Multiple calls are done in order to get more reliable time estimates. The number of calls is determined automatically, but you can use the -r and -n options to specify them directly. Type %timeit? to get more information.
Profiling Python code
The %timeit magic command gives you precious information about the total time taken by a function or a statement. This can help you find the fastest among several implementations of an algorithm, for example.
When you're finding that some code is too slow, you need to profile it before you can make it faster. Profiling gives you more than the total time taken by a function; it tells you exactly what is taking too long in your code.
The %prun magic command lets you easily profile your code. It provides a convenient interface to Python's native profile module.
Let's see a simple example. We first create a function returning the number of connected components in a file, as follows:
In [44]: import networkx In [45]: def ncomponents(file): graph = networkx.read_edgelist(file) return networkx.number_connected_components(graph)
Now we write a function that returns the number of connected components in all graphs defined in the directory, as follows:
In [46]: import glob def ncomponents_files(): return [(file, ncomponents(file)) for file in sorted(glob.glob('*.edges'))]
The glob module (https://docs.python.org/3.4/library/glob.html) lets us find all files matching a given pattern (here, all files with the .edges file extension).
In [47]: for file, n in ncomponents_files(): print(file.ljust(12), n, 'component(s)') Out[47]: 0.edges 5 component(s) 107.edges 1 component(s) 1684.edges 4 component(s) 1912.edges 2 component(s) 3437.edges 2 component(s) 348.edges 1 component(s) 3980.edges 4 component(s) 414.edges 2 component(s) 686.edges 1 component(s) 698.edges 3 component(s)
Let's first evaluate the time taken by this function:
In [48]: %timeit ncomponents_files() Out[48]: 1 loops, best of 3: 634 ms per loop
Now, to run the profiler, we use the %prun magic function, as follows:
In [49]: %prun -s cumtime ncomponents_files() Out[49]: 2391070 function calls in 1.038 seconds Ordered by: cumulative time ncalls tottime percall cumtime percall filename:lineno(function) 1 0.000 0.000 1.038 1.038 {built-in method exec} 1 0.000 0.000 1.038 1.038 <string>:1(<module>) 10 0.000 0.000 0.995 0.100 <string>:1(read_edgelist) 10 0.000 0.000 0.995 0.100 decorators.py:155(_open_file) 10 0.376 0.038 0.995 0.099 edgelist.py:174(parse_edgelist) 170174 0.279 0.000 0.350 0.000 graph.py:648(add_edge) 170184 0.059 0.000 0.095 0.000 edgelist.py:366(<genexpr>) 10 0.000 0.000 0.021 0.002 connected.py:98(number_connected_components) 35 0.001 0.000 0.021 0.001 connected.py:22(connected_components)
Let's explain what happened here. The profiler kept track of all function calls (including functions internal to NetworkX and Python) performed while our ncomponents_files() function was running. There were 2,391,070 function calls. That's a lot! Opening a file, reading and parsing every line, creating the graphs, finding the number of connected components, and so on, are operations that involve many function calls.
The profiler shows the list of all function calls (we just showed a subset here). There are many ways to sort the functions. Here, we chose to sort them by cumulative time, which is the total time spent within every function (-s cumtime option).
For every function, the profiler shows the total number of calls, and several time statistics, described here (copied verbatim from the profiler documentation):
tottime: the total time spent in the given function (and excluding time made in calls to sub-functions)percall: the quotient oftottimedivided byncallscumtime: the cumulative time spent in this and all subfunctionspercall: the quotient ofcumtimedivided by the number of non-recursive function calls
You will find more information by typing %prun? or by looking here: https://docs.python.org/3.4/library/profile.html
Here, we see that computing the number of connected components took considerably less time than loading the graphs from the text files. Depending on the use-case, this might suggest using a more efficient file format.
There is of course much more to say about profiling and optimization. For example, it is possible to profile a function line by line, which provides an even more fine-grained profiling report. The IPython Cookbook contains many more details.






















































