The tidyverse is, according to its homepage, "an opinionated collection of R packages designed for data science" (see https://www.tidyverse.org/). All of the packages are installed using install.packages("tidyverse"), and calling library(tidyverse) loads a subset of these packages, those considered to have the most value in day-to-day data science. Calling library(tidyverse) loads the following packages:
- ggplot2: For plotting
- dplyr: For data wrangling
- tidyr: For tidying (and untidying!) data
- readr: Better functions for reading comma- and tab-delimited data and other types of flat files
- purrr: For iterating
- tibble: Better dataframes
- stringr: For dealing with strings
- forcats: Better handling of factors, an R property used to describe the categories of a variable, described briefly earlier in the chapter
Installing the tidyverse also installs the following packages, which then need to be loaded separately with their own library() instruction: readxl, haven, jsonlite, xml2, httr, rvest, DBI, lubridate, hms, blob, rlang, magrittr, glue, and broom. For more details on these packages, consult the documentation at tidyverse.org/.
The key word in this description is opinionated. The tidyverse is a set of R packages that work with tidy data, either producing it, or consuming it, or both. Tidy data was described by Hadley Wickham in the Journal of Statistical Software, Vol 59, Issue 10 (https://www.jstatsoft.org/article/view/v059i10/v59i10.pdf). Tidy data obeys three principles:
- Each variable forms a column
- Each observation forms a row
- Each type of observational unit forms a table
For many R users, their first introduction to tidy data will have been ggplot2, which consumes tidy data and requires other types of data to be munged into this form. In order to explain what this means, we will look at a simple example. For the purposes of this discussion, we will ignore the last principle, which is more about organizing groups of datasets rather than individual datasets.
Let's have a look at a simple example of a messy dataset. In the real world, you will find datasets that are a lot messier than this, but this will serve to illustrate the principles we are using here. The following are the first three rows of the medal table for the Pyeongchang Winter Olympics, which took place in 2018, as an R dataframe:
medals = data.frame(country = c("Norway", "Germany", "Canada"),
gold = c(14, 14, 11),
silver = c(14, 10, 8),
bronze = c(11, 7, 10)
)
If we print it at the console, it looks like the following screenshot:
This is perhaps the most common sort of messy data you will come across: great as a summary, certainly intelligible to people watching the Winter Olympics, but not tidy. There are medals in three different columns. A tidy dataset would contain only one column for the medal tallies. Let's tidy it up using the tidyr package, which is loaded with library(tidyverse), or you can load it separately with library(tidyr). We can tidy the data very simply using the gather() function. The gather() function takes a dataframe as an argument, along with key and value column names (which you can set to what you like), and the column names that you wish to be gathered (all other columns will be duplicated as appropriate). In this case, we want to gather everything except the country, so we can use -variableName to indicate that we wish to gather everything except variableName. The final code looks like the following:
library(tidyr)
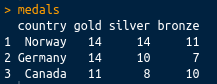
gather(medals, key = Type, value = Medals, -country)
This has a nice tidy output, as shown in the following screenshot: